Difference between revisions of "Flattop k-eff sensitivity example"
(Add serpentTools examples for visualizing sensitivity profiles) |
Ana Jambrina (Talk | contribs) (→Input) |
||
| Line 58: | Line 58: | ||
% --- Use unresolved resonance sampling for the fast system | % --- Use unresolved resonance sampling for the fast system | ||
| − | set ures 1 | + | set ures 1 |
% --- Define Vitamin-J energy grid (one of the defaults) | % --- Define Vitamin-J energy grid (one of the defaults) | ||
Latest revision as of 15:54, 5 June 2022
This example demonstrates some basic sensitivity calculation capabilities for the Popsy (Flattop) critical experiment (PU-MET-FAST-006 in the ICSBEP handbook).
Contents
Description
The experimental configuration consists of a plutonium alloy sphere (radius 4.5332 cm) reflected by a natural uranium blanket (outer radius 24.1420 cm). The multiplication factor of the system is close to unity (being a critical experiment). This input calculates the sensitivity of the multiplication factor to perturbations applied to the cross sections of 235U, 238U, 239Pu and 240Pu. A processing script for MATLAB/OCTAVE is provided for printing and plotting out the sensitivities.
Input
Input for Popsy (Flattop) sensitivity calculation example:
% --- Core outer surface and blanket outer surface surf s1 sph 0.0 0.0 0.0 4.5332 surf s2 sph 0.0 0.0 0.0 24.1420 % --- Divide space to core, blanket and outside cell c1 0 core -s1 cell c3 0 blanket s1 -s2 cell c4 0 outside s2 % --- Core is made from plutonium alloy mat core sum 94239.03c 3.6697e-2 94240.03c 1.8700e-3 94241.03c 1.1639e-4 31000.03c 1.4755e-3 % --- Blanket is made from natural uranium mat blanket sum 92234.03c 2.6438e-6 92235.03c 3.4610e-4 92238.03c 4.7721e-2 % --- Cross section libraries %set acelib "<path-to-acelib>" % --- Neutron population set pop 10000 100000 50 1 100 100 % --- Use a central point source as the initial source src point sp 0.0 0.0 0.0 % --- Write output each 100 cycles set outp 100 % --- Modify event buffer size to 3 events per neutron set nbuf 3 3 % --- Use unresolved resonance sampling for the fast system set ures 1 % --- Define Vitamin-J energy grid (one of the defaults) ene myvit-j 4 nj23 % %%% % %%% --- Sensitivity options % %%% % --- Run sensitivity calculation using Vitamin-J energy grid sens opt egrid myvit-j % --- Use 15 latent generations for the sensitivity calculations sens opt latgen 15 % --- Use 10 generations for Iterated Fission Probability set ifp 10 % %%% % %%% --- Sensitivity responses % %%% % --- Calculate sensitivity of k-effective to perturbations sens resp keff % %%% % %%% --- Sensitivity perturbations % %%% % --- Separate perturbations for the different nuclides and total sens pert zailist 922350 922380 942390 942400 total % --- Do not calculate material-wise perturbations (only total) sens pert matlist total % --- Perturb cross sections separately for each sum reaction mode (fission, capture etc.) sens pert xs all % --- Score sensitivity results directly to a matrix for each particle % Assume 0.2 matrices needed per generation per particle sens opt direct 0.2
Output
The contents of some long arrays have been replaced with "...".
Output for Popsy (Flattop) sensitivity calculation example:
% Number of different bins in sensitivity calculation: SENS_N_MAT = 1; SENS_N_ZAI = 5; SENS_N_PERT = 7; SENS_N_ENE = 175; % Materials included in sensitivity calculation: SENS_MAT_LIST = [ 'total ' ]; % Indices for different materials: iSENS_MAT_TOT = 1; % Nuclides included in sensitivity calculation: SENS_ZAI_LIST = [ 0 % total 922350 922380 942390 942400 ]; % Indices for different ZAIs: iSENS_ZAI_TOT = 1; iSENS_ZAI_922350 = 2; iSENS_ZAI_922380 = 3; iSENS_ZAI_942390 = 4; iSENS_ZAI_942400 = 5; % Reactions included in sensitivity calculation: SENS_PERT_LIST = [ 'total xs ' 'ela scatt xs ' 'sab scatt xs ' 'inl scatt xs ' 'capture xs ' 'fission xs ' 'nxn xs ' ]; % Indices for different perturbations: iSENS_PERT_TOT_XS = 1; iSENS_PERT_ELA_XS = 2; iSENS_PERT_SAB_XS = 3; iSENS_PERT_INL_XS = 4; iSENS_PERT_CAPT_XS = 5; iSENS_PERT_FISS_XS = 6; iSENS_PERT_NXN_XS = 7; % Sensitivity calculation energy group boundaries: SENS_E = [ ... ]; % Sensitivity calculation energy group lethargy widths: SENS_LETHARGY_WIDTHS = [ ... ]; % Sensitivities with 15 latent generations: % Effective multiplication factor: ADJ_PERT_KEFF_SENS = [ ... ]; ADJ_PERT_KEFF_SENS = reshape(ADJ_PERT_KEFF_SENS, [2, SENS_N_ENE, SENS_N_PERT, SENS_N_ZAI, SENS_N_MAT]); ADJ_PERT_KEFF_SENS = permute(ADJ_PERT_KEFF_SENS, [5, 4, 3, 2, 1]); ADJ_PERT_KEFF_SENS_E_INT = [ ... ]; ADJ_PERT_KEFF_SENS_E_INT = reshape(ADJ_PERT_KEFF_SENS_E_INT, [2, SENS_N_PERT, SENS_N_ZAI, SENS_N_MAT]); ADJ_PERT_KEFF_SENS_E_INT = permute(ADJ_PERT_KEFF_SENS_E_INT, [4, 3, 2, 1]);
Data-processing and plotting
Script for printing and plotting the results tested with MATLAB R2014a and OCTAVE 4.0.0:
%% --- Flag to print out zero values
printzeros = false;
%% --- Input file name
fname = 'input';
%% --- Run the sensitivity output file
run([fname '_sens0.m']);
%% --- Print some basic output data
disp(sprintf('Results from Serpent sensitivity calculation using:\n'));
%% --- Materials
disp([num2str(SENS_N_MAT) ' perturbed materials:']);
for i=1:1:SENS_N_MAT
disp(SENS_MAT_LIST(i,:));
end
%% --- Nuclides
disp(sprintf(['\n' num2str(SENS_N_ZAI) ' perturbed nuclides (ZAIs):']));
for i=1:1:SENS_N_ZAI
disp(sprintf('%d',SENS_ZAI_LIST(i,:)));
end
%% --- Perturbations
disp(sprintf(['\n' num2str(SENS_N_PERT) ' different perturbations:']));
for i=1:1:SENS_N_PERT
disp(SENS_PERT_LIST(i,:));
end
%% --- Energy grid
disp(sprintf(['\nEnergy grid with ' num2str(SENS_N_ENE) ' bins.']));
%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
%% -- Output sensitivities one by one %%%%%%%%%%%%%%%%%%%%%%%%%
%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
disp(sprintf('\n********************\n\nSensitivity results:\n'))
txt = sprintf('%-20s %-6s %-20s : %12s | 2 sigma confidence interval', 'Material', 'ZAI', ' Perturbation', 'sensitivity');
disp(txt)
%% --- This script only prints/plots the k-eff sensitivities
%% You can change the following three lines to plot something else
res = ADJ_PERT_KEFF_SENS;
resE = ADJ_PERT_KEFF_SENS_E_INT;
resStr = 'K-eff';
for iMat=1:1:SENS_N_MAT
matStr = SENS_MAT_LIST(iMat,:);
for iZai=1:1:SENS_N_ZAI
myZai = SENS_ZAI_LIST(iZai);
%% --- Turn ZAI into a string for printing
if (myZai == 0)
zaiStr = sprintf('%-6s ', 'total');
else
zaiStr = sprintf('%6d ', SENS_ZAI_LIST(iZai));
end
%% --- Last loop is over perturbations
for iPert=1:1:SENS_N_PERT
%% --- Turn perturbation into a string for printing
pertStr = sprintf('%-20s ', SENS_PERT_LIST(iPert,:));
val = resE(iMat,iZai,iPert,1);
err = resE(iMat,iZai,iPert,2);
%% --- Calculate 2 sigma confidence interval assuming
%% normally distributed random variable
minV = min([val*(1-2*err), val*(1+2*err)]);
maxV = max([val*(1-2*err), val*(1+2*err)]);
%% --- Create a formatted string for printing out the sensitivity
txt = sprintf('%-20s %-6s %-20s: %+12.8f | [%+12.8f, %+12.8f]', matStr, zaiStr, pertStr, val, minV, maxV);
%% --- Print formatted string
if (printzeros || (val ~= 0) || (err ~= 0))
disp(txt);
%% --- Move on to plotting a stairstep plot
%% --- Convert energy grid to eV
xvals = SENS_E*1e6;
%% --- Get energy dependent sensitivity and relative error
yvals = squeeze(res(iMat, iZai, iPert, :, 1))./SENS_LETHARGY_WIDTHS';
yerr = squeeze(res(iMat, iZai, iPert, :, 2));
%% --- We'll need to re-append the last value if we want to
%% get the stairstep plot correct
yvals = [yvals; yvals(end)];
yerr = [yerr; yerr(end)];
%% --- Create the figure for plotting
fig1 = figure('visible', 'off');
%% --- Plot the sample mean
h1 = stairs(xvals, yvals, 'k-');
set(h1, 'LineWidth', 2);
%% --- Plot the sample mean +- 2*sqrt(sample variance)
hold on;
h2 = stairs(xvals, yvals.*(1+2*yerr), 'r-');
h3 = stairs(xvals, yvals.*(1-2*yerr), 'r-');
%% --- Fill the area of non-zero-sensitivity
N = size(xvals,2)*2-2;
xx = zeros(N,1);
yy = zeros(N,1);
xx(1:2:N) = xvals(1:1:end-1);
xx(2:2:N) = xvals(2:1:end);
yy(1:2:N) = yvals(1:1:end-1);
yy(2:2:N) = yvals(1:1:end-1);
h4 = fill(xx, yy, [1.0 0.8 0.8]);
hold off;
%% --- Make the picture a bit nicer
set(gca,'xscale','log');
set(gca,'XTick', [1e4 1e5 1e6 1e7]);
grid on;
box on;
xlim([1e4, SENS_E(end)*1e6]);
ylim([-0.1, 0.5]);
%% --- Label the axes and add a title
xlabel('Energy (eV)');
ylabel('Sensitivity per unit lethargy');
txt = sprintf('Response: %s\nMaterial: %s\nZAI: %s\n perturbation %s', resStr, matStr, zaiStr, pertStr);
title(txt);
%% --- Save the plot to file
filename = sprintf('%s_sens_%ld_%ld_%ld.png', resStr, iMat, iZai, iPert);
print(filename,'-dpng');
%% --- Close figure
close(fig1);
end
end
%% --- Add empty line after each ZAI
disp(sprintf(' '));
end
end
Scripts for visualizing with Python
The following two scripts rely on the serpentTools python package for the parsing of the output and plotting https://serpent-tools.readthedocs.io/en/latest/
Fonts are configured with matplotlib to look like LaTeX https://matplotlib.org/tutorials/introductory/customizing.html
Full example of the sensitivity reader can be found at https://serpent-tools.readthedocs.io/en/latest/examples/Sensitivity.html
Simple python processing script:
"""Script for visualizing k-eff sensitivities for Flattop problem"""
from matplotlib import pyplot
import serpentTools
s = serpentTools.read("flattop_sens.m")
# Only response is required, but then it plots against all
# permutations of materials, perturbations, and materials
# Pass material and/or isotope ZAI and/or specific perturbation to plot
# sensitivities due to those properties only
# s.plot handles log scaling, step plotting, and uncertainty
s.plot(
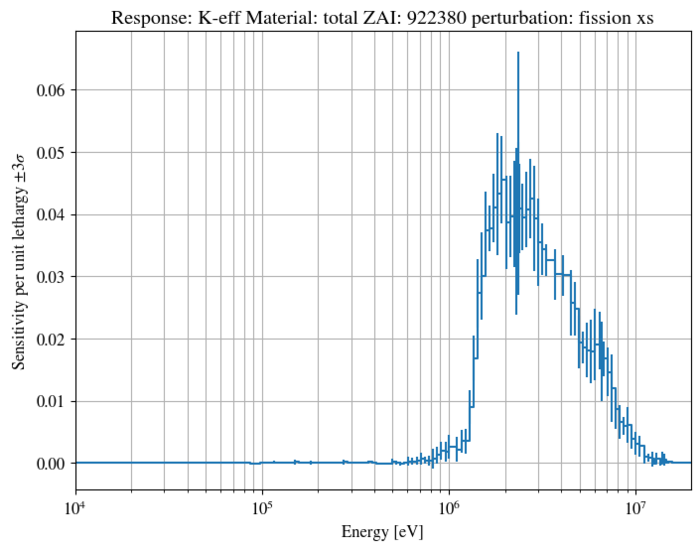
"keff", mat="total", zai=922380, pert="fission xs",
)
# Format the plot to look like the MATLAB / OCTAVE example
# No y axis limits
pyplot.xlim(1E4, s.energies[-1]*1E6)
# Turn on minor grids for the x-axis to show the log scale better
pyplot.grid("x", which="both")
pyplot.grid("y")
# Add a title to the top of the figure
pyplot.title("Response: K-eff Material: total ZAI: 922380 perturbation: fission xs")
# Show the figure
pyplot.show()
Detailed python processing script:
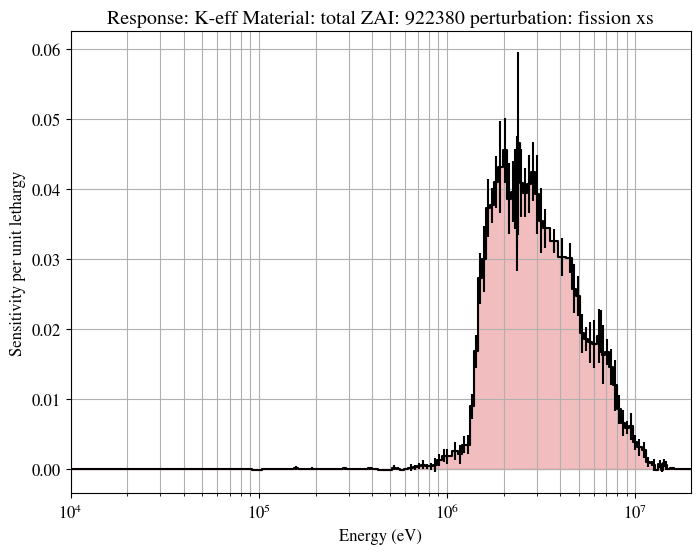
This script seeks to fully replicate the figures provided in the Wiki made using the MATLAB / OCTAVE programs
from matplotlib import pyplot
import serpentTools
s = serpentTools.read("flattop_sens.m")
# Extract the response of interest from the sensitivities dictionary
ks = s.sensitivities["keff"]
# Obtain the expected value and uncertainty by indexing into the multi-dimensional
# response. The matrix is structured [materials, isotopes, perturbations, energies, 2]
# where the last dimension stores value and uncertainty
# We can use the materials, zais, and perts dictionaries to obtain the correct index
# in each dimension for the quantity of interest
kslice = ks[
s.materials["total"], # index for sensitivity due to all materials
s.zais[922380], # index for sensitivity due to U238
s.perts["fission xs"], # index for sensitivity due to fission xs
]
# Normalize per unit lethargy
# Python arrays are zero-indexed, so the expected value is in the 0 index
value = kslice[:, 0] / s.lethargyWidths
# Compute 2-sigma uncertainty
unc = kslice[:, 1] * 2 * value
# Draw errorbars
# The energy vector has one additional entry, so we will instead drop the first item
# by slicing from the second position forward
pyplot.errorbar(
s.energies[1:] * 1E6, # convert to MeV
value, # expected value
yerr=unc, # uncertainty
drawstyle="steps-mid", # step-plot
c="k", # draw using a black line
)
# Shade the region under the curve red
# Use a mild transparency
pyplot.fill_between(s.energies[1:]*1E6, value, color="tab:red", alpha=0.3, step="mid")
# Format the plot
pyplot.xscale("log")
pyplot.xlim(1E4, s.energies[-1]*1E6)
# Major and minor grids for the log-scaled x axis
pyplot.grid("x", which="both")
pyplot.grid("y")
pyplot.xlabel("Energy (eV)")
pyplot.ylabel("Sensitivity per unit lethargy")
pyplot.title("Response: K-eff Material: total ZAI: 922380 perturbation: fission xs")
pyplot.show()
Results
Runtime of the input on a computer cluster node with a Intel Xeon E5-2690 v2 processor (10 physical cores, 20 logical cores) using 20 OpenMP threads took approximately 4 hours 50 minutes and used (allocated) 6117 MB of RAM. The run produced an 133 KB [input]_sens.m file.
Executing the processing and plotting script produced the following output using OCTAVE 4.0.0
Output from the processing script:
Results from Serpent sensitivity calculation using: 1 perturbed materials: total 5 perturbed nuclides (ZAIs): 0 922350 922380 942390 942400 7 different perturbations: total xs ela scatt xs sab scatt xs inl scatt xs capture xs fission xs nxn xs Energy grid with 175 bins. ******************** Sensitivity results: Material ZAI Perturbation : sensitivity | 2 sigma confidence interval total total total xs : +0.90527000 | [ +0.90411125, +0.90642875] total total ela scatt xs : +0.16188300 | [ +0.16084695, +0.16291905] total total inl scatt xs : +0.07863120 | [ +0.07815941, +0.07910299] total total capture xs : -0.05447680 | [ -0.05459665, -0.05435695] total total fission xs : +0.71923200 | [ +0.71897308, +0.71949092] total total nxn xs : +0.00125871 | [ +0.00122598, +0.00129144] total 922350 total xs : +0.00818980 | [ +0.00809971, +0.00827989] total 922350 ela scatt xs : +0.00081898 | [ +0.00074855, +0.00088941] total 922350 inl scatt xs : +0.00038832 | [ +0.00035803, +0.00041861] total 922350 capture xs : -0.00054016 | [ -0.00054913, -0.00053120] total 922350 fission xs : +0.00752267 | [ +0.00747753, +0.00756781] total 922350 nxn xs : +0.00000661 | [ +0.00000463, +0.00000859] total 922380 total xs : +0.21901900 | [ +0.21805532, +0.21998268] total 922380 ela scatt xs : +0.13649800 | [ +0.13562441, +0.13737159] total 922380 inl scatt xs : +0.06499990 | [ +0.06459690, +0.06540290] total 922380 capture xs : -0.03974930 | [ -0.03986060, -0.03963800] total 922380 fission xs : +0.05727050 | [ +0.05714450, +0.05739650] total 922380 nxn xs : +0.00096484 | [ +0.00093783, +0.00099186] total 942390 total xs : +0.65488800 | [ +0.65424621, +0.65552979] total 942390 ela scatt xs : +0.02212830 | [ +0.02159722, +0.02265938] total 942390 inl scatt xs : +0.01241460 | [ +0.01216631, +0.01266289] total 942390 capture xs : -0.01294550 | [ -0.01299469, -0.01289631] total 942390 fission xs : +0.63329000 | [ +0.63301135, +0.63356865] total 942390 nxn xs : +0.00027191 | [ +0.00025614, +0.00028768] total 942400 total xs : +0.02006720 | [ +0.01991469, +0.02021971] total 942400 ela scatt xs : +0.00129135 | [ +0.00117513, +0.00140757] total 942400 inl scatt xs : +0.00063176 | [ +0.00056606, +0.00069746] total 942400 capture xs : -0.00102246 | [ -0.00103534, -0.00100958] total 942400 fission xs : +0.01916650 | [ +0.01909367, +0.01923933] total 942400 nxn xs : +0.00000945 | [ +0.00000567, +0.00001323]
These results were obtained using ENDF/B-VII.1 based cross section libraries.
A number of produced figures are also shown here for ease of comparison: